Quantitative Methods
Experimental methods and tools, mathematical methods in biology, bioinformatics.
Looking for a broader view? This category is part of:
Experimental methods and tools, mathematical methods in biology, bioinformatics.
Looking for a broader view? This category is part of:
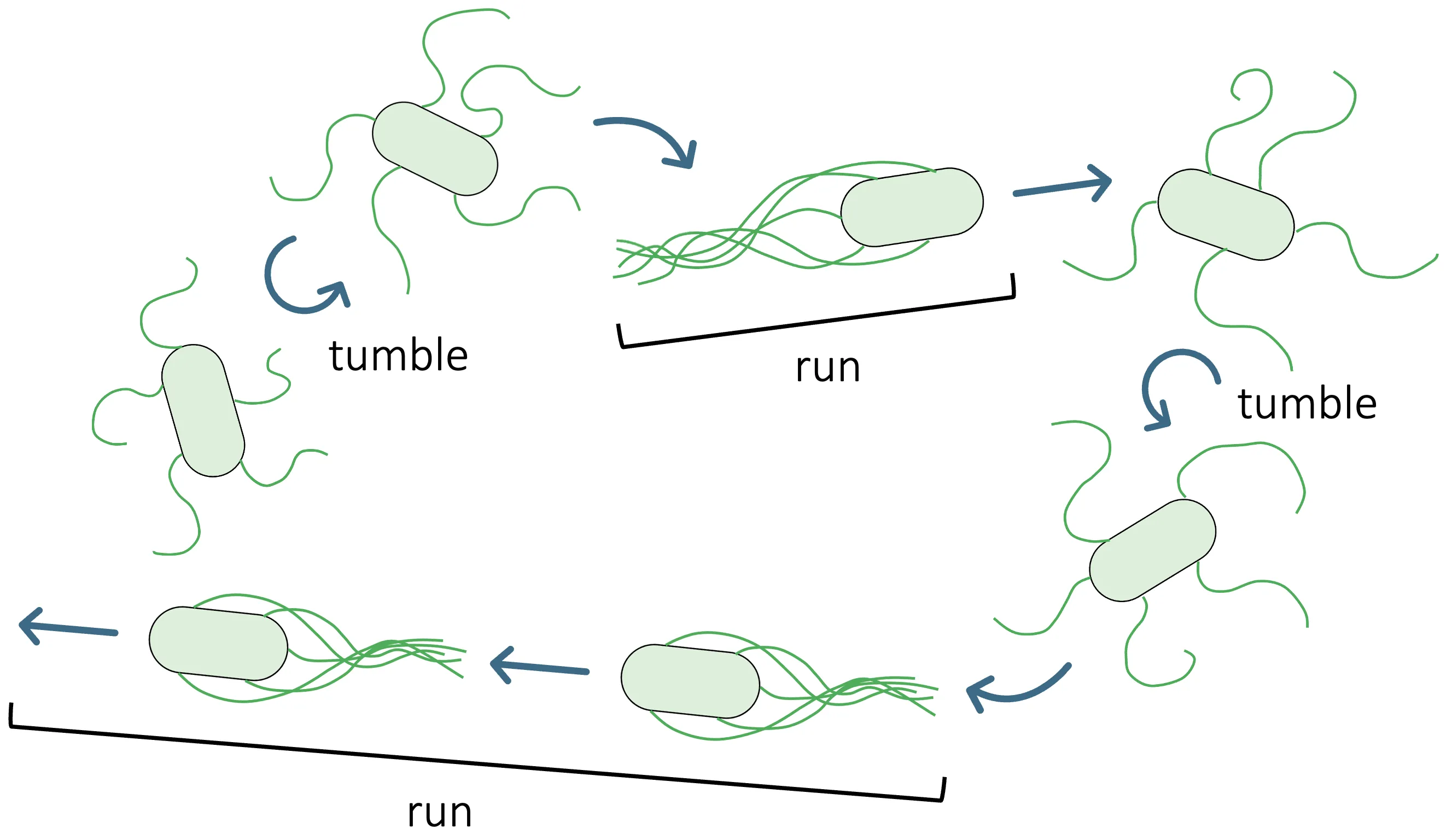
Bacterial chemotactic sensing converts noisy chemical signals into running and tumbling. We analyze the static sensing limits of mixed Tar/Tsr chemoreceptor clusters in individual Escherichia coli cells using a heterogeneous Monod-Wyman-Changeux (MWC) model. By sweeping a seven-dimensional parameter space, we compute three sensing performance metrics-channel capacity, effective Hill coefficient, and dynamic range. Across E. coli-like parameter regimes, we consistently observe pronounced local maxima of channel capacity, whereas neither the effective Hill coefficient nor the dynamic range exhibit comparable optimization. The capacity-achieving input distribution is bimodal, which implies that individual cells maximize information by sampling both low- and high concentration regimes. Together, these results suggest that, at the individual-cell level, channel capacity may be selected for in E. coli receptor clusters.
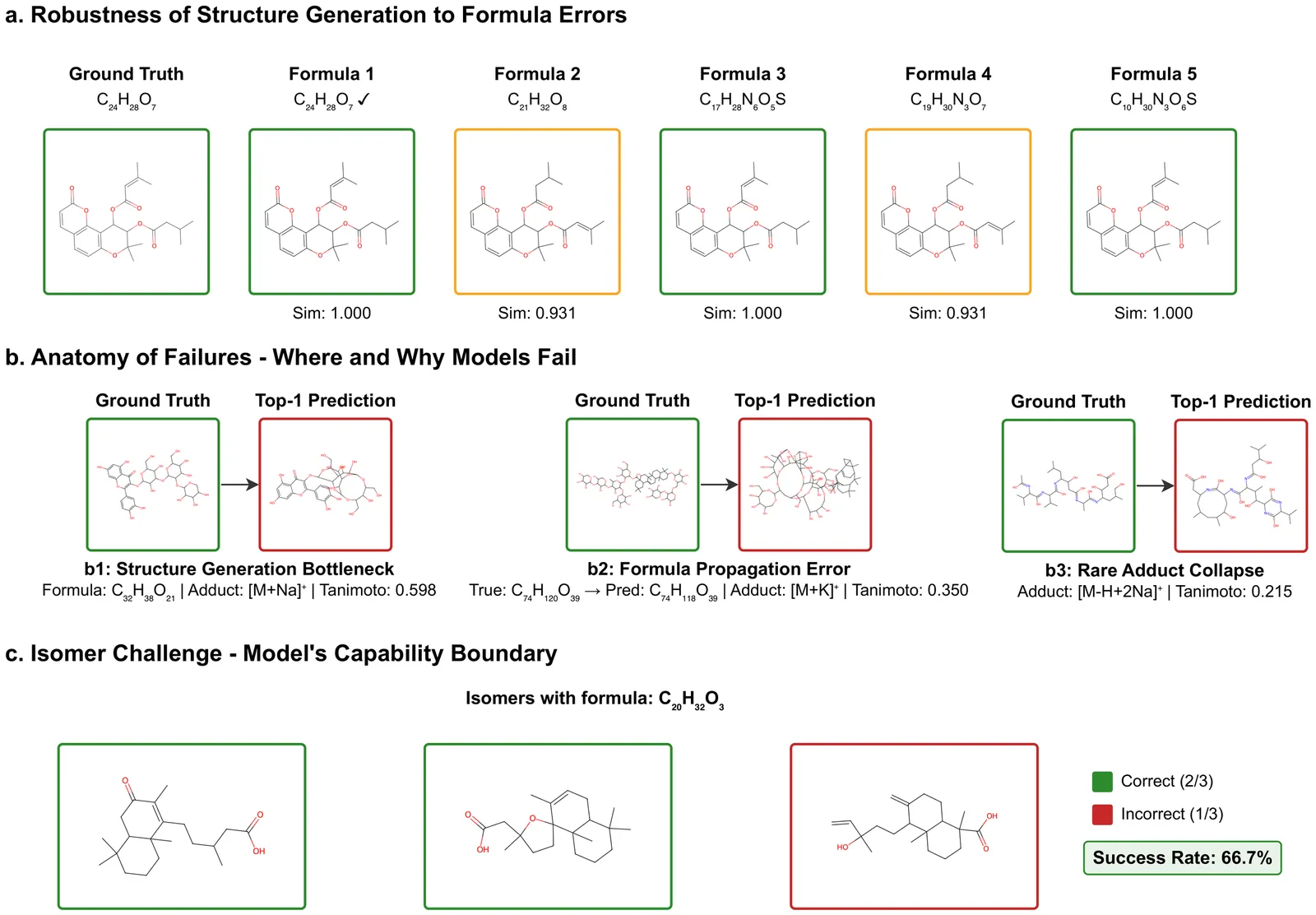
Liquid chromatography mass spectrometry (LC-MS)-based metabolomics and exposomics aim to measure detectable small molecules in biological samples. The results facilitate hypothesis-generating discovery of metabolic changes and disease mechanisms and provide information about environmental exposures and their effects on human health. Metabolomics and exposomics are made possible by the high resolving power of LC and high mass measurement accuracy of MS. However, a majority of the signals from such studies still cannot be identified or annotated using conventional library searching because existing spectral libraries are far from covering the vast chemical space captured by LC-MS/MS. To address this challenge and unleash the full potential of metabolomics and exposomics, a number of computational approaches have been developed to predict compounds based on tandem mass spectra. Published assessment of these approaches used different datasets and evaluation. To select prediction workflows for practical applications and identify areas for further improvements, we have carried out a systematic evaluation of the state-of-the-art prediction algorithms. Specifically, the accuracy of formula prediction and structure prediction was evaluated for different types of adducts. The resulting findings have established realistic performance baselines, identified critical bottlenecks, and provided guidance to further improve compound predictions based on MS.
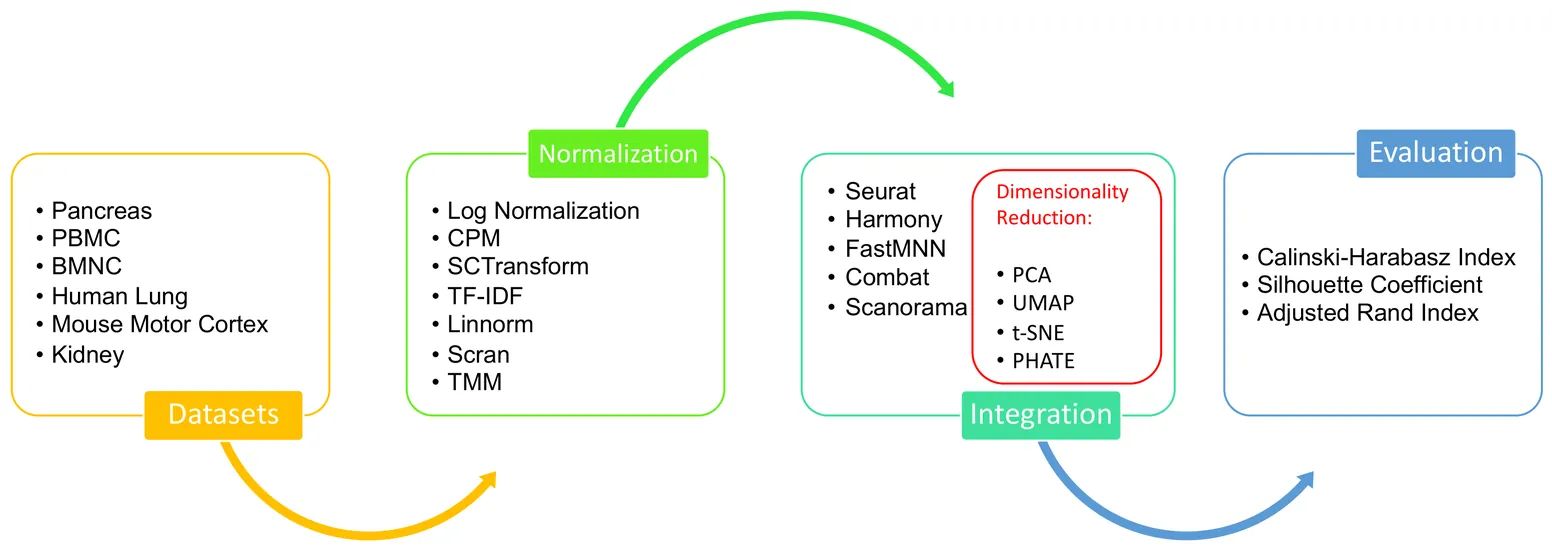
Single-cell data analysis has the potential to revolutionize personalized medicine by characterizing disease-associated molecular changes at the single-cell level. Advanced single-cell multimodal assays can now simultaneously measure various molecules (e.g., DNA, RNA, Protein) across hundreds of thousands of individual cells, providing a comprehensive molecular readout. A significant analytical challenge is integrating single-cell measurements across different modalities. Various methods have been developed to address this challenge, but there has been no systematic evaluation of these techniques with different preprocessing strategies. This study examines a general pipeline for single-cell data analysis, which includes normalization, data integration, and dimensionality reduction. The performance of different algorithm combinations often depends on the dataset sizes and characteristics. We evaluate six datasets across diverse modalities, tissues, and organisms using three metrics: Silhouette Coefficient Score, Adjusted Rand Index, and Calinski-Harabasz Index. Our experiments involve combinations of seven normalization methods, four dimensional reduction methods, and five integration methods. The results show that Seurat and Harmony excel in data integration, with Harmony being more time-efficient, especially for large datasets. UMAP is the most compatible dimensionality reduction method with the integration techniques, and the choice of normalization method varies depending on the integration method used.
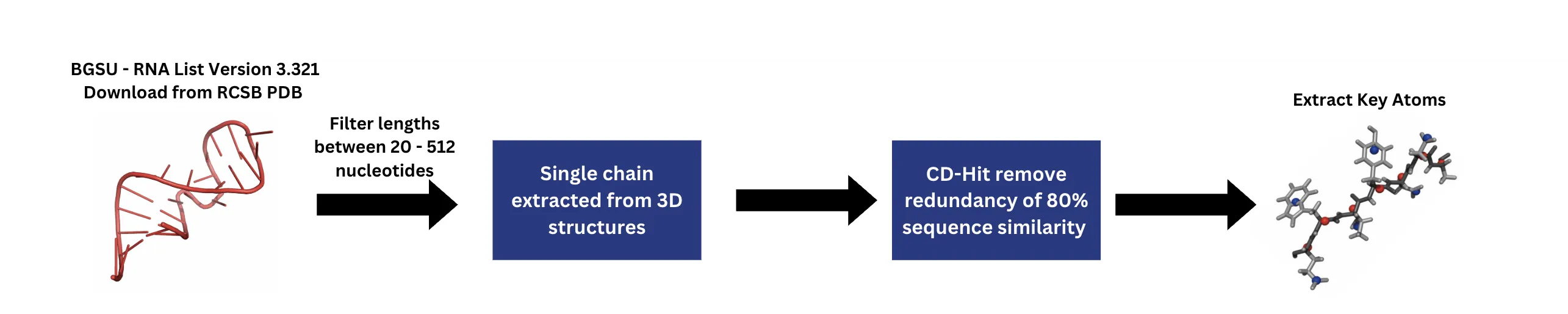
RNA's diverse biological functions stem from its structural versatility, yet accurately predicting and designing RNA sequences given a 3D conformation (inverse folding) remains a challenge. Here, I introduce a deep learning framework that integrates Geometric Vector Perceptron (GVP) layers with a Transformer architecture to enable end-to-end RNA design. I construct a dataset consisting of experimentally solved RNA 3D structures, filtered and deduplicated from the BGSU RNA list, and evaluate performance using both sequence recovery rate and TM-score to assess sequence and structural fidelity, respectively. On standard benchmarks and RNA-Puzzles, my model achieves state-of-the-art performance, with recovery and TM-scores of 0.481 and 0.332, surpassing existing methods across diverse RNA families and length scales. Masked family-level validation using Rfam annotations confirms strong generalization beyond seen families. Furthermore, inverse-folded sequences, when refolded using AlphaFold3, closely resemble native structures, highlighting the critical role of geometric features captured by GVP layers in enhancing Transformer-based RNA design.
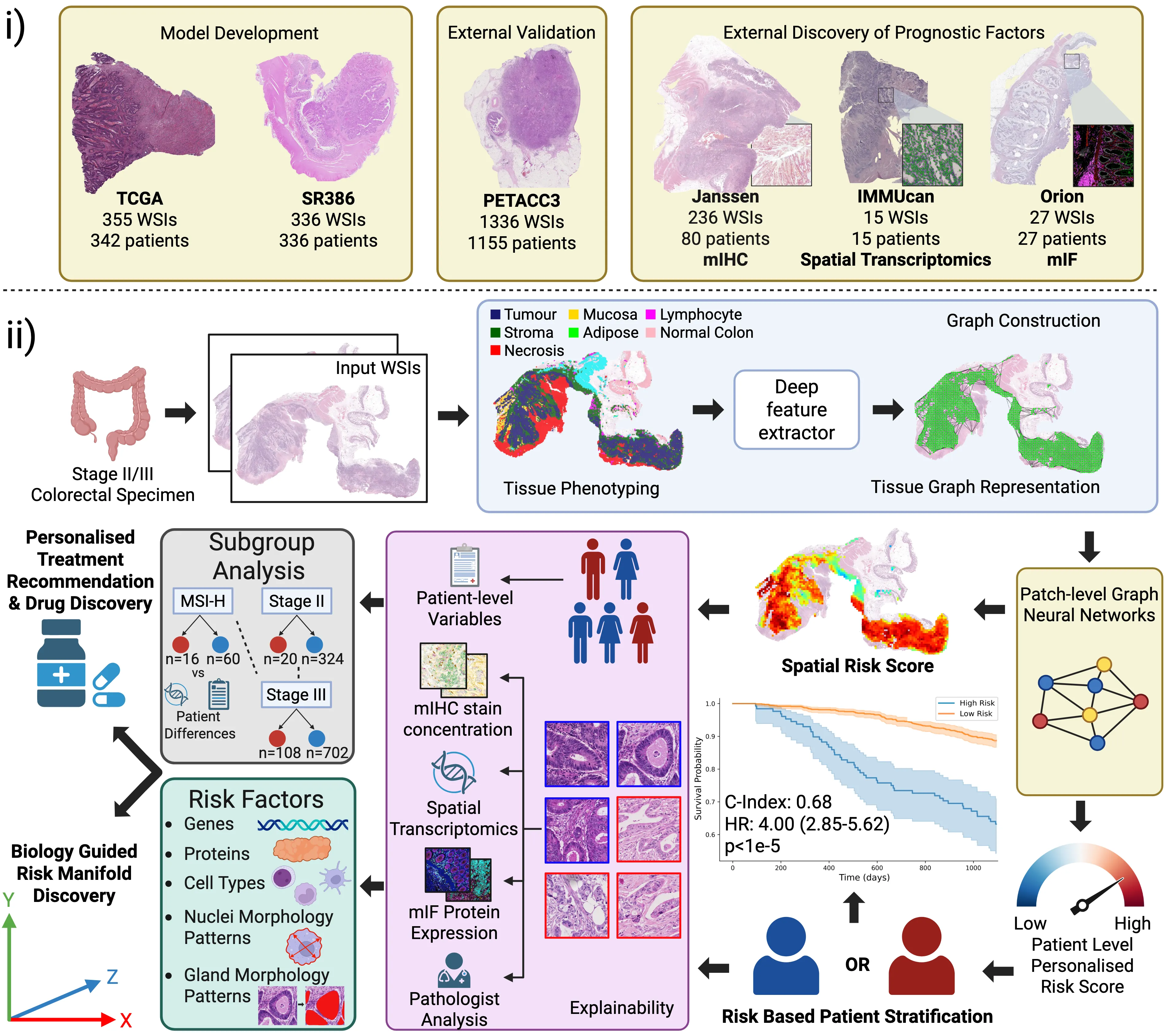
Routine histology contains rich prognostic information in stage II/III colorectal cancer, much of which is embedded in complex spatial tissue organisation. We present INSIGHT, a graph neural network that predicts survival directly from routine histology images. Trained and cross-validated on TCGA (n=342) and SURGEN (n=336), INSIGHT produces patient-level spatially resolved risk scores. Large independent validation showed superior prognostic performance compared with pTNM staging (C-index 0.68-0.69 vs 0.44-0.58). INSIGHT spatial risk maps recapitulated canonical prognostic histopathology and identified nuclear solidity and circularity as quantitative risk correlates. Integrating spatial risk with data-driven spatial transcriptomic signatures, spatial proteomics, bulk RNA-seq, and single-cell references revealed an epithelium-immune risk manifold capturing epithelial dedifferentiation and fetal programs, myeloid-driven stromal states including $\mathrm{SPP1}^{+}$ macrophages and $\mathrm{LAMP3}^{+}$ dendritic cells, and adaptive immune dysfunction. This analysis exposed patient-specific epithelial heterogeneity, stratification within MSI-High tumours, and high-risk routes of CDX2/HNF4A loss and CEACAM5/6-associated proliferative programs, highlighting coordinated therapeutic vulnerabilities.
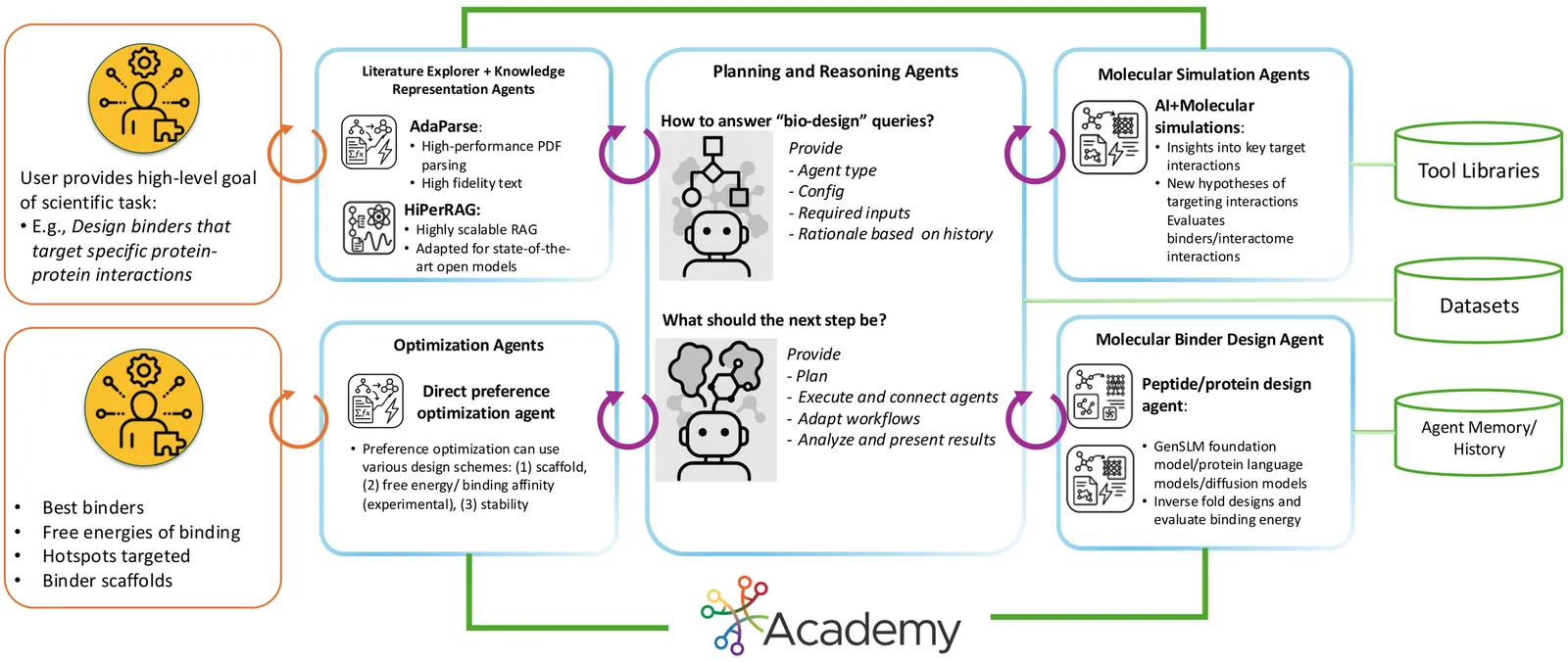
Intrinsically disordered proteins (IDPs) represent crucial therapeutic targets due to their significant role in disease -- approximately 80\% of cancer-related proteins contain long disordered regions -- but their lack of stable secondary/tertiary structures makes them "undruggable". While recent computational advances, such as diffusion models, can design high-affinity IDP binders, translating these to practical drug discovery requires autonomous systems capable of reasoning across complex conformational ensembles and orchestrating diverse computational tools at scale.To address this challenge, we designed and implemented StructBioReasoner, a scalable multi-agent system for designing biologics that can be used to target IDPs. StructBioReasoner employs a novel tournament-based reasoning framework where specialized agents compete to generate and refine therapeutic hypotheses, naturally distributing computational load for efficient exploration of the vast design space. Agents integrate domain knowledge with access to literature synthesis, AI-structure prediction, molecular simulations, and stability analysis, coordinating their execution on HPC infrastructure via an extensible federated agentic middleware, Academy. We benchmark StructBioReasoner across Der f 21 and NMNAT-2 and demonstrate that over 50\% of 787 designed and validated candidates for Der f 21 outperformed the human-designed reference binders from literature, in terms of improved binding free energy. For the more challenging NMNAT-2 protein, we identified three binding modes from 97,066 binders, including the well-studied NMNAT2:p53 interface. Thus, StructBioReasoner lays the groundwork for agentic reasoning systems for IDP therapeutic discovery on Exascale platforms.
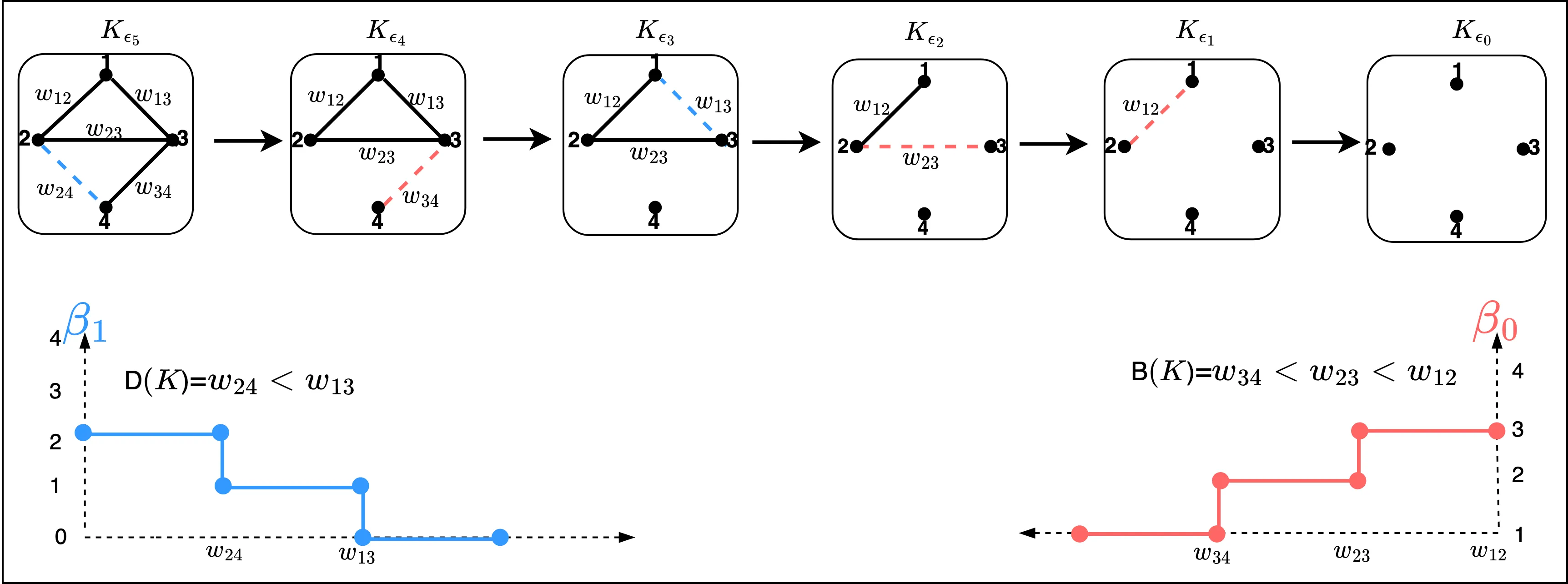
Identifying and comparing topological features, particularly cycles, across different topological objects remains a fundamental challenge in persistent homology and topological data analysis. This work introduces a novel framework for constructing cycle communities through two complementary approaches. First, a dendrogram-based methodology leverages merge-tree algorithms to construct hierarchical representations of homology classes from persistence intervals. The Wasserstein distance on merge trees is introduced as a metric for comparing dendrograms, establishing connections to hierarchical clustering frameworks. Through simulation studies, the discriminative power of dendrogram representations for identifying cycle communities is demonstrated. Second, an extension of Stratified Gradient Sampling simultaneously learns multiple filter functions that yield cycle barycenter functions capable of faithfully reconstructing distinct sets of cycles. The set of cycles each filter function can reconstruct constitutes cycle communities that are non-overlapping and partition the space of all cycles. Together, these approaches transform the problem of cycle matching into both a hierarchical clustering and topological optimization framework, providing principled methods to identify similar topological structures both within and across groups of topological objects.
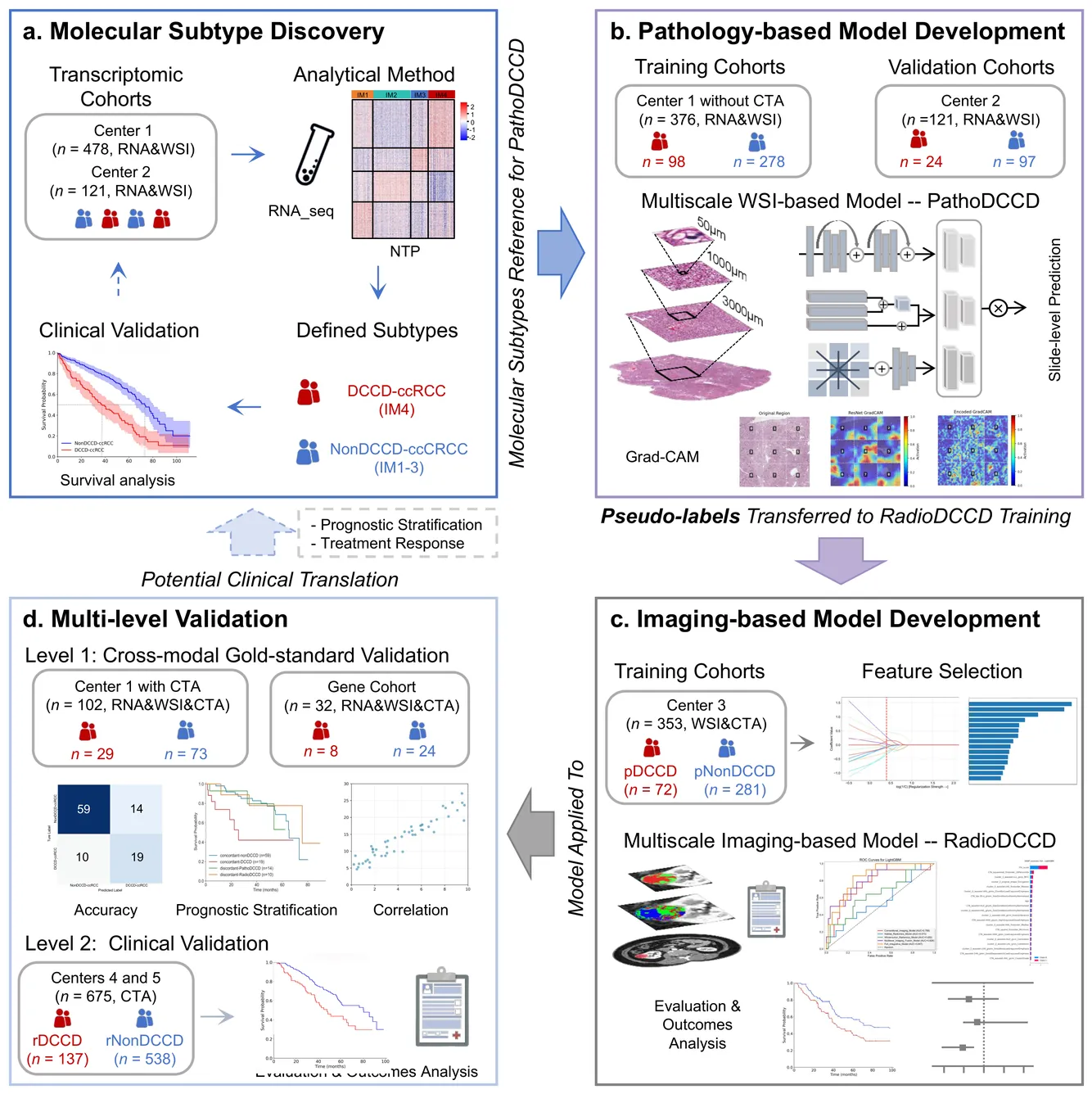
Clear cell renal cell carcinoma (ccRCC) exhibits extensive intratumoral heterogeneity on multiple biological scales, contributing to variable clinical outcomes and limiting the effectiveness of conventional TNM staging, which highlights the urgent need for multiscale integrative analytic frameworks. The lipid-deficient de-clear cell differentiated (DCCD) ccRCC subtype, defined by multi-omics analyses, is associated with adverse outcomes even in early-stage disease. Here, we establish a hierarchical cross-scale framework for the preoperative identification of DCCD-ccRCC. At the highest layer, cross-modal mapping transferred molecular signatures to histological and CT phenotypes, establishing a molecular-to-pathology-to-radiology supervisory bridge. Within this framework, each modality-specific model is designed to mirror the inherent hierarchical structure of tumor biology. PathoDCCD captured multi-scale microscopic features, from cellular morphology and tissue architecture to meso-regional organization. RadioDCCD integrated complementary macroscopic information by combining whole-tumor and its habitat-subregions radiomics with a 2D maximal-section heterogeneity metric. These nested models enabled integrated molecular subtype prediction and clinical risk stratification. Across five cohorts totaling 1,659 patients, PathoDCCD reliably recapitulated molecular subtypes, while RadioDCCD provided reliable preoperative prediction. The consistent predictions identified patients with the poorest clinical outcomes. This cross-scale paradigm unifies molecular biology, computational pathology, and quantitative radiology into a biologically grounded strategy for preoperative noninvasive molecular phenotyping of ccRCC.
 2512.12272
2512.12272Post-translational modifications (PTMs) serve as a dynamic chemical language regulating protein function, yet current proteomic methods remain blind to a vast portion of the modified proteome. Standard database search algorithms suffer from a combinatorial explosion of search spaces, limiting the identification of uncharacterized or complex modifications. Here we introduce OmniNovo, a unified deep learning framework for reference-free sequencing of unmodified and modified peptides directly from tandem mass spectra. Unlike existing tools restricted to specific modification types, OmniNovo learns universal fragmentation rules to decipher diverse PTMs within a single coherent model. By integrating a mass-constrained decoding algorithm with rigorous false discovery rate estimation, OmniNovo achieves state-of-the-art accuracy, identifying 51\% more peptides than standard approaches at a 1\% false discovery rate. Crucially, the model generalizes to biological sites unseen during training, illuminating the dark matter of the proteome and enabling unbiased comprehensive analysis of cellular regulation.
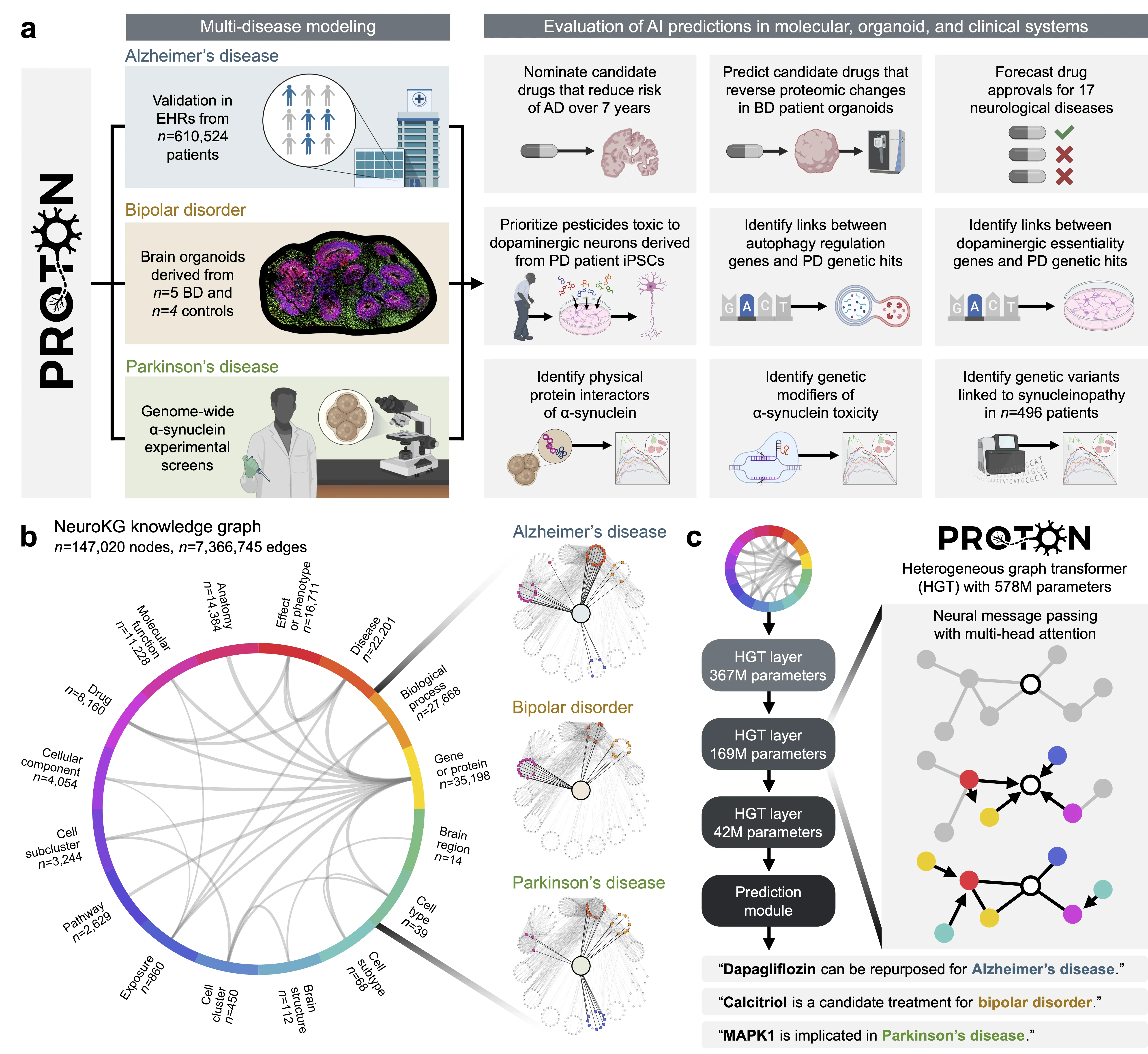
Neurological diseases are the leading global cause of disability, yet most lack disease-modifying treatments. We present PROTON, a heterogeneous graph transformer that generates testable hypotheses across molecular, organoid, and clinical systems. To evaluate PROTON, we apply it to Parkinson's disease (PD), bipolar disorder (BD), and Alzheimer's disease (AD). In PD, PROTON linked genetic risk loci to genes essential for dopaminergic neuron survival and predicted pesticides toxic to patient-derived neurons, including the insecticide endosulfan, which ranked within the top 1.29% of predictions. In silico screens performed by PROTON reproduced six genome-wide $α$-synuclein experiments, including a split-ubiquitin yeast two-hybrid system (normalized enrichment score [NES] = 2.30, FDR-adjusted $p < 1 \times 10^{-4}$), an ascorbate peroxidase proximity labeling assay (NES = 2.16, FDR $< 1 \times 10^{-4}$), and a high-depth targeted exome sequencing study in 496 synucleinopathy patients (NES = 2.13, FDR $< 1 \times 10^{-4}$). In BD, PROTON predicted calcitriol as a candidate drug that reversed proteomic alterations observed in cortical organoids derived from BD patients. In AD, we evaluated PROTON predictions in health records from $n = 610,524$ patients at Mass General Brigham, confirming that five PROTON-predicted drugs were associated with reduced seven-year dementia risk (minimum hazard ratio = 0.63, 95% CI: 0.53-0.75, $p < 1 \times 10^{-7}$). PROTON generated neurological hypotheses that were evaluated across molecular, organoid, and clinical systems, defining a path for AI-driven discovery in neurological disease.
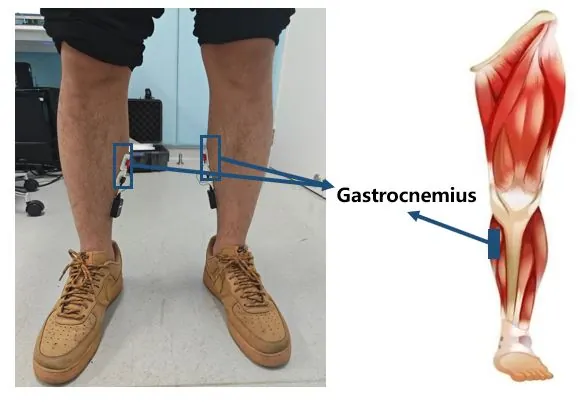
Rehabilitation exoskeletons have shown promising results in promoting recovery for stroke patients. Accurately and timely identifying the motion intentions of patients is a critical challenge in enhancing active participation during lower limb exoskeleton-assisted rehabilitation training. This paper proposes a Dual-Channel Attentive Fusion Network (DCAF-Net) that synergistically integrates pre-movement surface electromyography (sEMG) and inertial measurement unit (IMU) data for lower limb intention prediction in stroke patients. First, a dual-channel adaptive channel attention module is designed to extract discriminative features from 48 time-domain and frequency-domain features derived from bilateral gastrocnemius sEMG signals. Second, an IMU encoder combining convolutional neural network (CNN) and attention-based long short-term memory (attention-LSTM) layers is designed to decode temporal-spatial movement patterns. Third, the sEMG and IMU features are fused through concatenation to enable accurate recognition of motion intention. Extensive experiment on 11 participants (8 stroke subjects and 3 healthy subjects) demonstrate the effectiveness of DCAF-Net. It achieved a prediction accuracies of 97.19% for patients and 93.56% for healthy subjects. This study provides a viable solution for implementing intention-driven human-in-the-loop assistance control in clinical rehabilitation robotics.
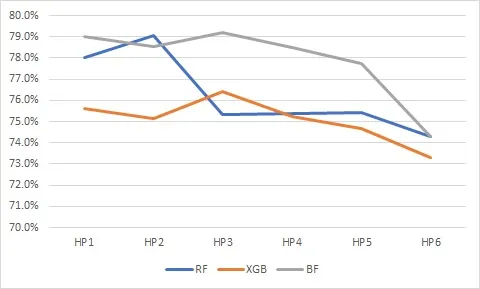
Cancer patients may undergo lengthy and painful chemotherapy treatments, comprising several successive regimens or plans. Treatment inefficacy and other adverse events can lead to discontinuation (or failure) of these plans, or prematurely changing them, which results in a significant amount of physical, financial, and emotional toxicity to the patients and their families. In this work, we build treatment failure models based on the Real World Evidence (RWE) gathered from patients' profiles available in our oncology EMR/EHR system. We also describe our feature engineering pipeline, experimental methods, and valuable insights obtained about treatment failures from trained models. We report our findings on five primary cancer types with the most frequent treatment failures (or discontinuations) to build unique and novel feature vectors from the clinical notes, diagnoses, and medications that are available in our oncology EMR. After following a novel three axes - performance, complexity and explainability - design exploration framework, boosted random forests are selected because they provide a baseline accuracy of 80% and an F1 score of 75%, with reduced model complexity, thus making them more interpretable to and usable by oncologists.
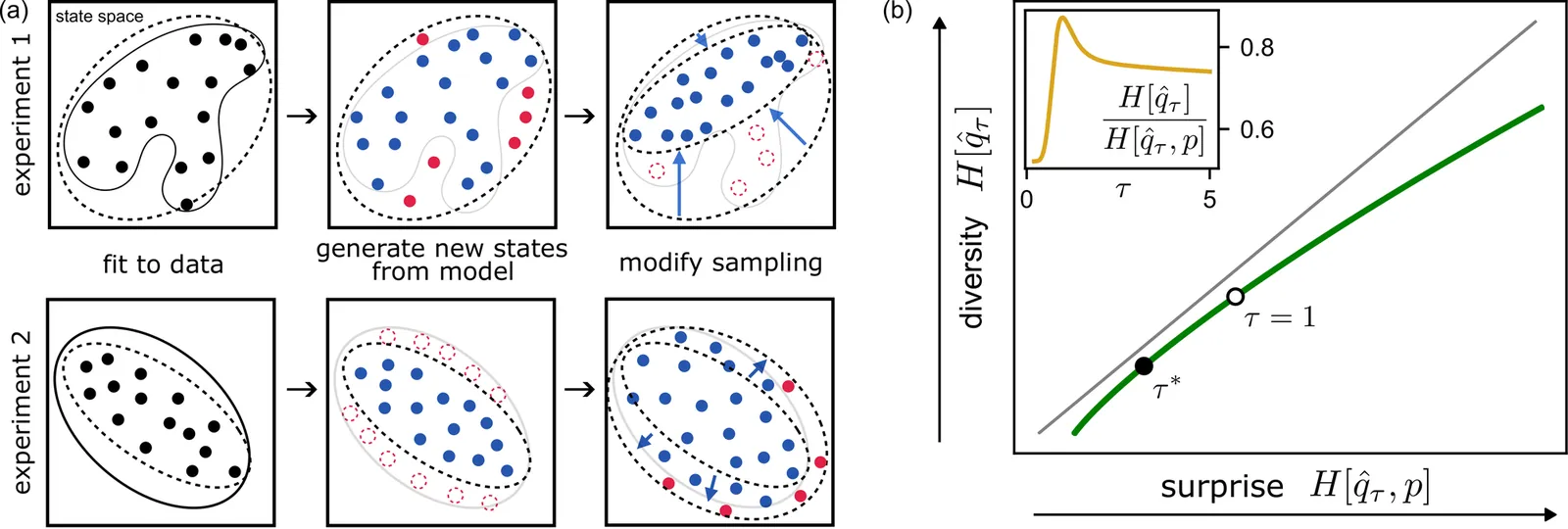
Generative models of complex systems often require post-hoc parameter adjustments to produce useful outputs. For example, energy-based models for protein design are sampled at an artificially low ''temperature'' to generate novel, functional sequences. This temperature tuning is a common yet poorly understood heuristic used across machine learning contexts to control the trade-off between generative fidelity and diversity. Here, we develop an interpretable, physically motivated framework to explain this phenomenon. We demonstrate that in systems with a large ''energy gap'' - separating a small fraction of meaningful states from a vast space of unrealistic states - learning from sparse data causes models to systematically overestimate high-energy state probabilities, a bias that lowering the sampling temperature corrects. More generally, we characterize how the optimal sampling temperature depends on the interplay between data size and the system's underlying energy landscape. Crucially, our results show that lowering the sampling temperature is not always desirable; we identify the conditions where \emph{raising} it results in better generative performance. Our framework thus casts post-hoc temperature tuning as a diagnostic tool that reveals properties of the true data distribution and the limits of the learned model.
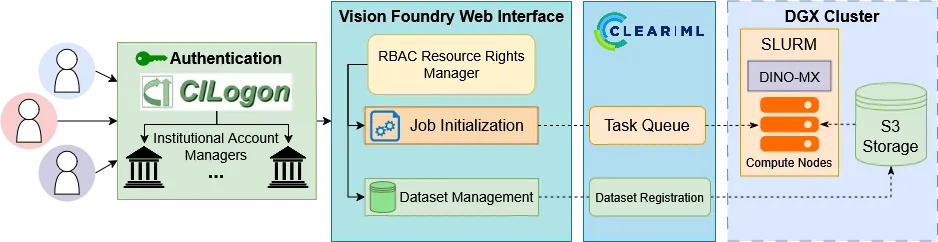
Self-supervised learning (SSL) leverages vast unannotated medical datasets, yet steep technical barriers limit adoption by clinical researchers. We introduce Vision Foundry, a code-free, HIPAA-compliant platform that democratizes pre-training, adaptation, and deployment of foundational vision models. The system integrates the DINO-MX framework, abstracting distributed infrastructure complexities while implementing specialized strategies like Magnification-Aware Distillation (MAD) and Parameter-Efficient Fine-Tuning (PEFT). We validate the platform across domains, including neuropathology segmentation, lung cellularity estimation, and coronary calcium scoring. Our experiments demonstrate that models trained via Vision Foundry significantly outperform generic baselines in segmentation fidelity and regression accuracy, while exhibiting robust zero-shot generalization across imaging protocols. By bridging the gap between advanced representation learning and practical application, Vision Foundry enables domain experts to develop state-of-the-art clinical AI tools with minimal annotation overhead, shifting focus from engineering optimization to clinical discovery.
In multicellular organisms, cells coordinate their activities through cell-cell communication (CCC), which are crucial for development, tissue homeostasis, and disease progression. Recent advances in single-cell and spatial omics technologies provide unprecedented opportunities to systematically infer and analyze CCC from these omics data, either by integrating prior knowledge of ligand-receptor interactions (LRIs) or through de novo approaches. A variety of computational methods have been developed, focusing on methodological innovations, accurate modeling of complex signaling mechanisms, and investigation of broader biological questions. These advances have greatly enhanced our ability to analyze CCC and generate biological hypotheses. Here, we introduce the biological mechanisms and modeling strategies of CCC, and provide a focused overview of more than 140 computational methods for inferring CCC from single-cell and spatial transcriptomic data, emphasizing the diversity in methodological frameworks and biological questions. Finally, we discuss the current challenges and future opportunities in this rapidly evolving field.
In cell culture bioprocessing, real-time batch process monitoring (BPM) refers to the continuous tracking and analysis of key process variables such as viable cell density, nutrient levels, metabolite concentrations, and product titer throughout the duration of a batch run. This enables early detection of deviations and supports timely control actions to ensure optimal cell growth and product quality. BPM plays a critical role in ensuring the quality and regulatory compliance of biopharmaceutical manufacturing processes. However, the development of accurate soft sensors for BPM is hindered by key challenges, including limited historical data, infrequent feedback, heterogeneous process conditions, and high-dimensional sensory inputs. This study presents a comprehensive benchmarking analysis of machine learning (ML) methods designed to address these challenges, with a focus on learning from historical data with limited volume and relevance in the context of bioprocess monitoring. We evaluate multiple ML approaches including feature dimensionality reduction, online learning, and just-in-time learning across three datasets, one in silico dataset and two real-world experimental datasets. Our findings highlight the importance of training strategies in handling limited data and feedback, with batch learning proving effective in homogeneous settings, while just-in-time learning and online learning demonstrate superior adaptability in cold-start scenarios. Additionally, we identify key meta-features, such as feed media composition and process control strategies, that significantly impact model transferability. The results also suggest that integrating Raman-based predictions with lagged offline measurements enhances monitoring accuracy, offering a promising direction for future bioprocess soft sensor development.
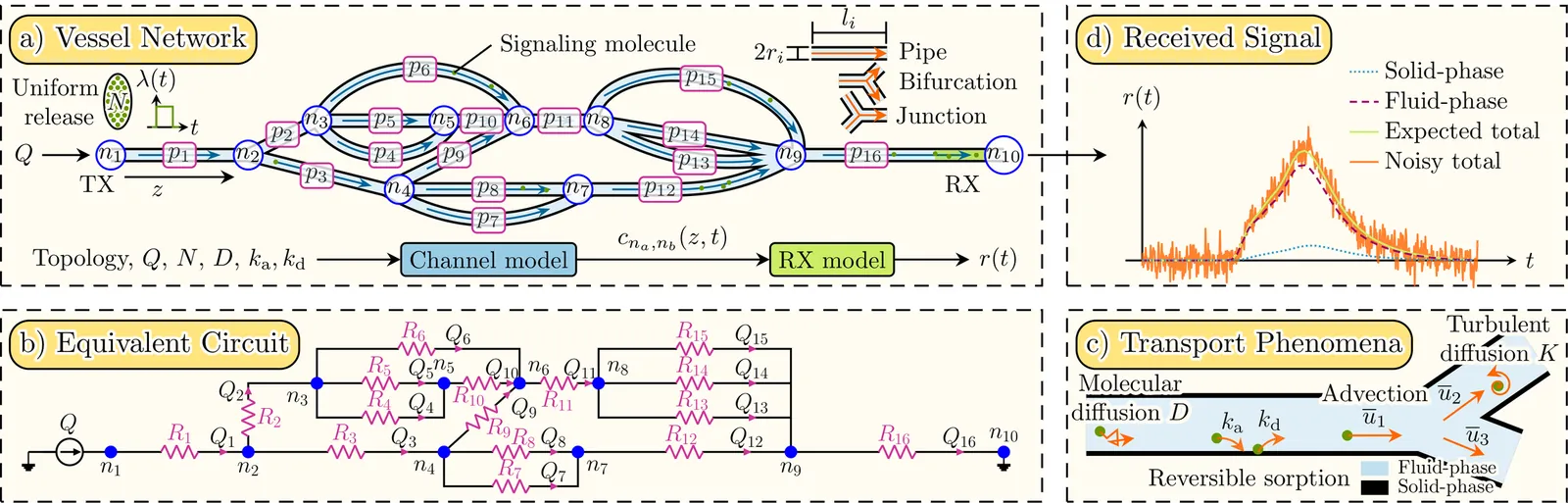
The notion of synthetic molecular communication (MC) refers to the transmission of information via signaling molecules and is foreseen to enable innovative medical applications in the human cardiovascular system (CVS). Crucially, the design of such applications requires accurate and experimentally validated channel models that characterize the propagation of signaling molecules, not just in individual blood vessels, but in complex vessel networks (VNs), as prevalent in the CVS. However, experimentally validated models for MC in VNs remain scarce. To address this gap, we propose a novel channel model for MC in complex VN topologies, which captures molecular transport via advection, molecular and turbulent diffusion, as well as adsorption and desorption at the vessel walls. We specialize this model for superparamagnetic iron-oxide nanoparticles (SPIONs) as signaling molecules by introducing a new receiver (RX) model for planar coil inductive sensors, enabling end-to-end experimental validation with a dedicated SPION testbed. Validation covers a range of channel topologies, from single-vessel topologies to branched VNs with multiple paths between transmitter (TX) and RX. Additionally, to quantify how the VN topology impacts signal quality, and inspired by multi-path propagation models in conventional wireless communications, we introduce two metrics, namely molecule delay and multi-path spread. We show that these metrics link the VN structure to molecule dispersion induced by the VN and mediately to the resulting signal-to-noise ratio (SNR) at the RX. The proposed VN structure-SNR link is validated experimentally, demonstrating that the proposed framework can support tasks such as optimal sensor placement in the CVS or the identification of suitable testbed topologies for specific SNR requirements. All experimental data are openly available on Zenodo.
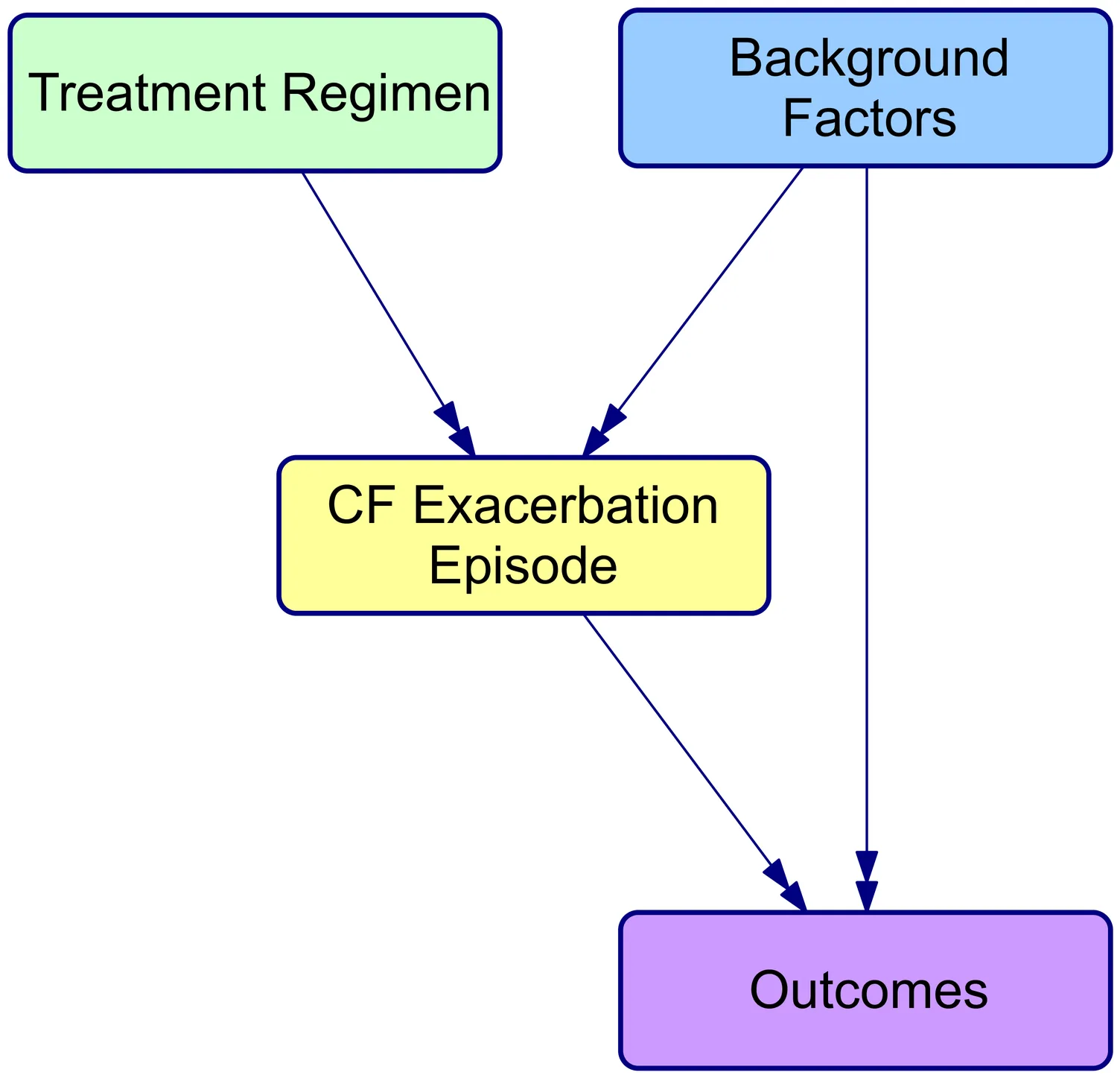
Loss of lung function in cystic fibrosis (CF) occurs progressively, punctuated by acute pulmonary exacerbations (PEx) in which abrupt declines in lung function are not fully recovered. A key component of CF management over the past half century has been the treatment of PEx to slow lung function decline. This has been credited with improvements in survival for people with CF (PwCF), but there is no consensus on the optimal approach to PEx management. BEAT-CF (Bayesian evidence-adaptive treatment of CF) was established to build an evidence-informed knowledge base for CF management. The BEAT-CF causal model is a directed acyclic graph (DAG) and Bayesian network (BN) for PEx that aims to inform the design and analysis of clinical trials comparing the effectiveness of alternative approaches to PEx management. The causal model describes relationships between background risk factors, treatments, and pathogen colonisation of the airways that affect the outcome of an individual PEx episode. The key factors, outcomes, and causal relationships were elicited from CF clinical experts and together represent current expert understanding of the pathophysiology of a PEx episode, guiding the design of data collection and studies and enabling causal inference. Here, we present the DAG that documents this understanding, along with the processes used in its development, providing transparency around our trial design and study processes, as well as a reusable framework for others.

Selecting an effective docking algorithm is highly context-dependent, and no single method performs reliably across structural, chemical, or protocol regimes. We introduce MolAS, a lightweight algorithm selection system that predicts per-algorithm performance from pretrained protein-ligand embeddings using attentional pooling and a shallow residual decoder. With only hundreds to a few thousand labelled complexes, MolAS achieves up to 15% absolute improvement over the single-best solver (SBS) and closes 17-66% of the Virtual Best Solver (VBS)-SBS gap across five diverse docking benchmarks. Analyses of reliability, embedding geometry, and solver-selection patterns show that MolAS succeeds when the oracle landscape exhibits low entropy and separable solver behaviour, but collapses under protocol-induced hierarchy shifts. These findings indicate that the main barrier to robust docking AS is not representational capacity but instability in solver rankings across pose-generation regimes, positioning MolAS as both a practical in-domain selector and a diagnostic tool for assessing when AS is feasible.
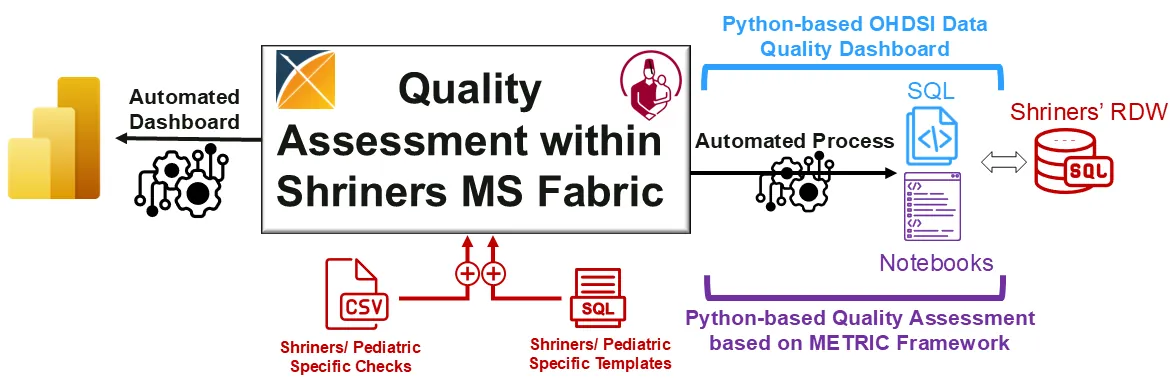
The rapid growth of Artificial Intelligence (AI) in healthcare has sparked interest in Trustworthy AI and AI Implementation Science, both of which are essential for accelerating clinical adoption. However, strict regulations, gaps between research and clinical settings, and challenges in evaluating AI systems continue to hinder real-world implementation. This study presents an AI implementation case study within Shriners Childrens (SC), a large multisite pediatric system, showcasing the modernization of SCs Research Data Warehouse (RDW) to OMOP CDM v5.4 within a secure Microsoft Fabric environment. We introduce a Python-based data quality assessment tool compatible with SCs infrastructure, extending OHDsi's R/Java-based Data Quality Dashboard (DQD) and integrating Trustworthy AI principles using the METRIC framework. This extension enhances data quality evaluation by addressing informative missingness, redundancy, timeliness, and distributional consistency. We also compare systematic and case-specific AI implementation strategies for Craniofacial Microsomia (CFM) using the FHIR standard. Our contributions include a real-world evaluation of AI implementations, integration of Trustworthy AI principles into data quality assessment, and insights into hybrid implementation strategies that blend systematic infrastructure with use-case-driven approaches to advance AI in healthcare.